Background
Several professors and students in the Biology and Computer Science departments are collecting E. Coli isolates and using a fast version of DNA sequencing called pyroprinting to create DNA fingerprints (called pyroprints) from them. These pyroprints are stored in a database application called CPLOP, the Cal Poly Library of Pyroprints. These isolates are clustered together using a combination of techniques such as hierarchical clustering to create strains.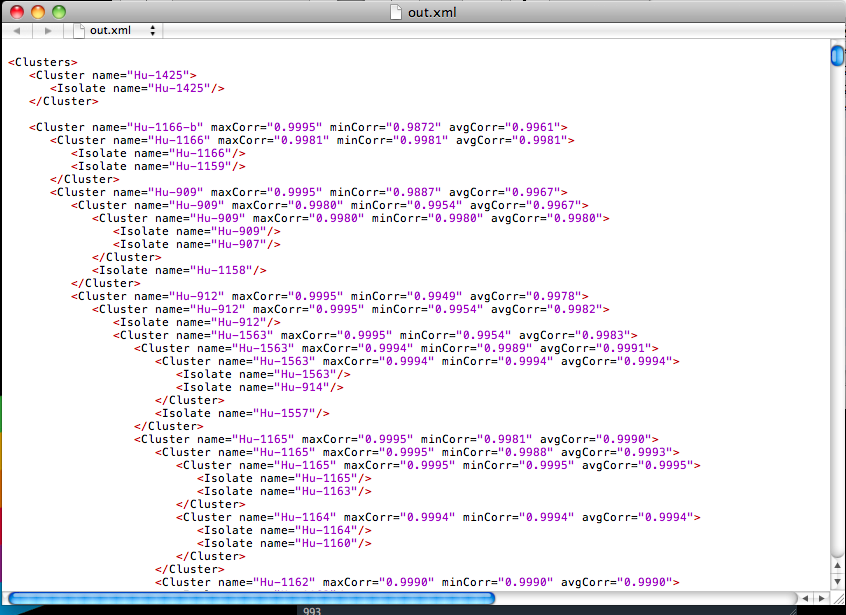
Problem
Currently, the clustering applications output the cluster information a XML and CSV files. Current researchers need a way to visualize clusters in order to derive important information about the clusters.The only way of measuring the similarity between clusters is by using Pearson correlation. This is our only dimension. Because of this limitation, we need to come up with new approaches to better present the data.
Implementation
This application was developed using Processing, an electronic sketchbook application developed at MIT.More information here.
Results
I implemented three visualizations that confirm characteristics about the clusters we have developed. The clustering algorithm was done on the 3,000 or so Isolates that we have stored in CPLOP.Linear Visualization
This was a simple visualization based on a level in a tree map. Each rectangle represents a node in a level of a tree. In this application, a user can click on a node to traverse the cluster hierarchy.
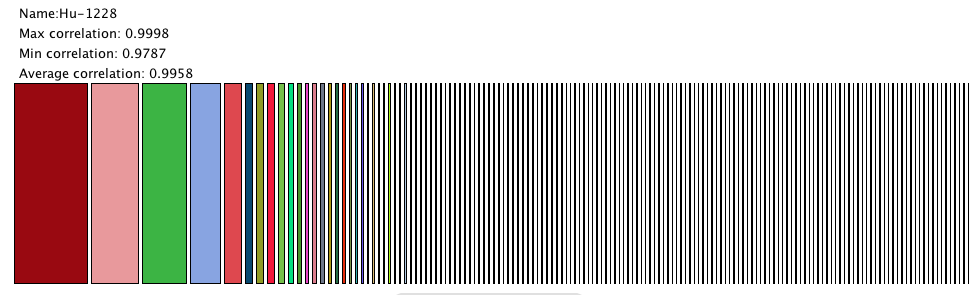
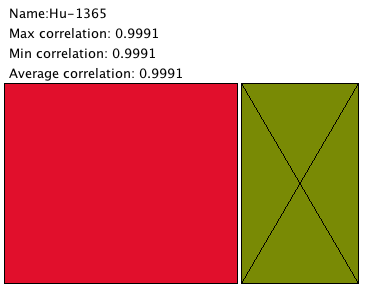
This visualization shows that our current data only has a few large clusters and lots of small ones. The small clusters are usually made up of one or individual isolates. While it's encouraging to see big clusters, the significant number of single-item clusters could mean there needs to be changes to our clustering algorithm.
The problem with this approach is that it does not use all of the white space on the screen. It will be more difficult to browse if the number of clusters increases in the future.
Spiral Visualization
To solve the problem of using space, I visualized the nodes as spheres along a spiral. This uses more space on the screen and looks more uniform between width and height than the linear visualization.
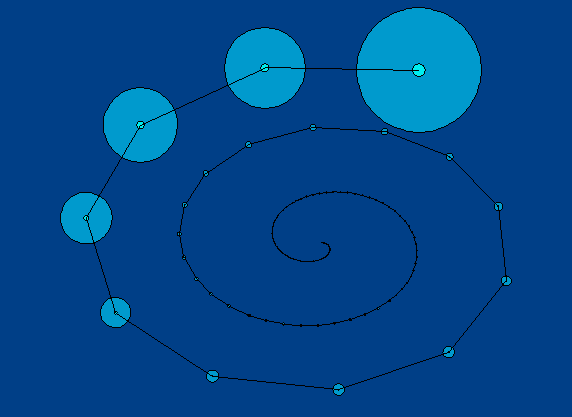
This visualization shows the few large clusters and numerous small clusters that the previous visualization. The scale of each cluster circle is proportional to the number of elements in that cluster.
This visualization presents other problems as well. I cannot easily show other information about the clusters, such as how wide they are.
8-Direction Visualization
Our last visualization aims to show multiple things. First we aim to show the number of elements in each cluster through scale as we did before. We also want to show the width of each element to show how items in a cluster vary between each other.
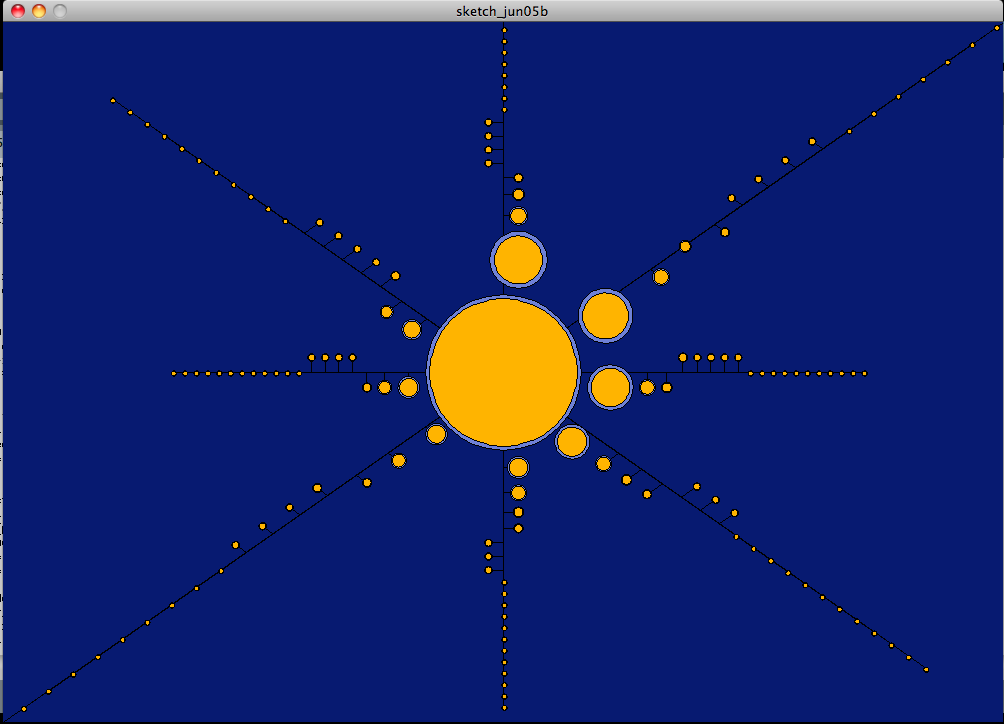
I scaled the circles for each cluster based on the number of items inside. I then placed them in order from largest to smallest counter-clockwise at 8 different directions. The clusters are then offset from the directional line based on their width. If the cluster's width is greater than the average, the cluster is offset to the right or bottom of the line. If the width is less than the average, then the cluster is offset to the left or top of the direction line.
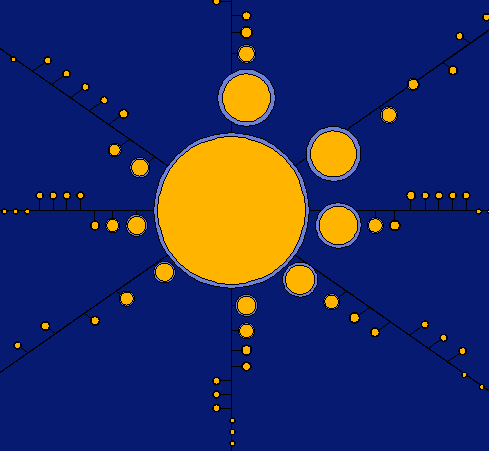

This visualization shows that the smaller clusters tend to hit around the average. Whereas the larger clusters tend to have a larger width.